The UIP pattern consists of ordinary locations of lung juxtaposed with areas of leukocyte infiltration and other locations with superior fibrosis in a given very low energy field. This variegated pattern has led for the hypothesis of temporal heterogeneity. This hypothesis states that UIP is definitely the con sequence of multiple episodes of irritation and fibrosis in response to multiple insults in excess of time, rather than the response to a single insult. Another histological fea ture of UIP is the presence of fibroblastic foci, parts of polyclonal fibroblast and myofibroblast aggregation and web site of deposition of new collagen, believed to signify areas of fibroblast proliferation and differentiation, and therefore are positioned on the edges of parts of dense fibrosis.
Provided their pivotal function while in the generation with the extracel lular matrix, fibroblasts and myofibroblasts are thought of the primary effector cells during the evolution of pulmonary fibrosis, inhibitor CX-4945 and their cellular supply is consequently a crucial query during the pathogenesis of fibrotic lung disorders. At the moment, 3 hypotheses deal with the origin of lung fibroblasts, The classical idea is the fact that tissue injury induces the activation, proliferation and differentiation of a resident fibroblast in the lung interstitial compartment into a myo fibroblast that migrates to the alveolar compartment and expresses constituents of the extracellular matrix leading to lung fibrosis. The second mechanism requires injury induced modifications from the microenvironment of the epithe lium or endothelium, inducing their transition to a mesenchymal phenotype and subsequently contributes to fibroproliferation.
The third mechanism pertains to a cir culating bone marrow derived progenitor cell, the fibro cyte, which can property to websites of lung injury, differentiate into fibroblasts and myofibroblasts, proliferate, and contri bute on the generation of extracellular matrix. Relevance of fibrocytes to lung fibrosis in animal models Making use of the mouse model of inhibitor PHA-665752 bleomycin administration, CD45 Col1 CXCR4 fibrocytes had been observed to increase inside the peripheral blood of animals chal lenged with intrapulmonary bleomycin as compared to saline, then return to steady state amounts. In con trast, lung fibrocytes started to appear in the lung 2 days after bleomycin administration, peak at 8 days immediately after bleo mycin and remained elevated in between days 16 and 20, this time program correlated temporally with deposition of col lagen while in the lungs and suggested mobi lization of fibrocytes in the bone marrow to your blood and subsequently on the lungs.
To tackle this difficulty defi nitively, a number of investigators have steady radiation chimeric mice through which bone marrow cells express enhanced GFP to the beta actin promoter. These will work have demon strated that in response to intra pulmonary bleomycin or lung irradiation, fibrocytes site visitors and accumulate within the lungs and also a subset differentiates into myelofibro blasts.
Monthly Archives: May 2014
We would recom mend iBMA dimension when gene precise external i
We would recom mend iBMA size when gene particular external informa tion is just not readily available. In Table two and Extra file 2, Table S1, each of the iBMA networks have been thresholded at a posterior probability of 50%. We located that iBMA prior also out performed other meth ods for these data over various posterior probability thresholds. Evaluation, transcription issue binding web site analysis In another assessment, we checked irrespective of whether the set our method to static information by getting rid of the subscript referring to your time stage from Equation, of target genes containing regarded binding internet sites for any certain TF were enriched between the kid nodes of your JASPAR database. Employing TFMscan, we retrieved a set of genes containing the recognized binding sites inside their upstream regions for each TF.
We then checked for enrichment of those genes among the inferred kid nodes of the corresponding TFs in each network with Fishers actual check. Table 3 reviews the quantity of TFs whose inferred selelck kinase inhibitor child nodes exhibited this kind of enrichment, at a false discovery rate of 10%. Each of the procedures that produced use of external information outperformed all of those who did not, illustrating the benefit of incorporating external expertise.LASSO shortlist and LAR shortlist appeared to produce somewhat superior benefits than iBMA prior on this binding web page analysis, nevertheless it is very likely the consequence of their more substantial network sizes. Comparison with Lirnet Lee et al. proposed a regression based mostly network development approach identified as Lirnet, which performed effectively on the publicly readily available gene expression information set from Brem et al.
The Brem data set recorded the steady state expression ranges for 112 selleck chemicals yeast segre gants, 95 of which were profiled in our time series experiments below different development conditions. Lee et al. showed that Lirnet out carried out Bayes ian networks over the similar data, and so we compared our best performer, iBMA prior, with Lirnet. Simply because Lirnet was formulated to analyze regular state ex pression information without time elements, we adapted We applied iBMA prior to precisely the same 3152 gene subset from the Brem et al. data that Lee et al. utilised. Lirnet constrained the search of regulators for every target gene to 304 known TFs. For fair com parison, we also confined the set of candidate regu lators to the similar TFs. Networks constructed from steady state gene expression information are unable to have feed back loops.
To detect and get rid of this kind of loops from our inferred network, we identified all strongly connected components employing the igraph R package,  and deleted the TF gene hyperlink associated with all the lowest posterior probability for every cycle. Similar as prior to, we evaluated distinctive procedures by assessing the concordance on the inferred networks together with the Yeastract database utilizing Pearsons chi square test. The evaluation resultin Table four demonstrate that iBMA prior outperformed Lirnet, nearly doubling the TPR and also the O/E ratio when making a comparable quantity of misclassified regulatory relationships. s
and deleted the TF gene hyperlink associated with all the lowest posterior probability for every cycle. Similar as prior to, we evaluated distinctive procedures by assessing the concordance on the inferred networks together with the Yeastract database utilizing Pearsons chi square test. The evaluation resultin Table four demonstrate that iBMA prior outperformed Lirnet, nearly doubling the TPR and also the O/E ratio when making a comparable quantity of misclassified regulatory relationships. s
16S rRNA gene sequence analyses Sequences from Bacteroidetes, sul
16S rRNA gene sequence analyses Sequences from Bacteroidetes, sulfate cutting down and sulfur oxidizing bacteria obtained from a preceding research had been employed to create phylo genetic trees. Briefly, 16S rRNA gene primers 8F and 787R were used to generate local community PCR goods, which were then cloned making use of TOPO TA vectors. Clones had been sequenced in each directions and assembled working with Sequencher computer software. Sequences have been assigned to distinct bacterial groups employing MOTHUR v1. 19. 2 with 97% sequence identity since the lower off stage for every Operational Taxonomic Unit. Phylogenetic trees have been constructed in the alignments primarily based around the Greatest Likelihood process and calculated utilizing Tamura Nei model. MEGA v5.03 was employed to build trees making use of 100 replicates to develop bootstrap self-assurance values. The Classifier instrument in the Ribosomal Database Venture II release ten. 26 and BLASTn had been used to classify and determine the nearest neighbors.
Cluster examination of wastewater concrete biofilms Cluster analysis based mostly around the transformed relative abundance data was utilised to review communi ties associated with various wastewater concrete bio films. Initial, we estimated the taxonomic distribution with the genus level of each and every microbial local community from selleck 16S rRNA gene pyrosequences generated in this review and Sanger chemistry 16S rRNA gene sequences produced in past scientific studies. This facts was applied to create Bray Curtis similarity coefficients in the trans formed information using the computer software Past v2. 03. This estimator compares the structures by accounting to the abundance distributions of attributes. Den drograms indicating romantic relationship of biofilms created by evaluating similarity coefficients estimates among sample web sites were calculated applying the UPGMA strategy with all the software MEGA v5. 03.
Metagenomic research Pyrosequencing was carried out applying the 454 Life Sciences GS FLX TitaniumW platform. Before sequence examination we implemented a dereplication pipeline to determine and eliminate clusters of artificially replicated sequences, i. e. reads that purchase Barasertib started at the exact same place but varied in length or con tained a sequencing discrepancy. Filter parameters integrated a cutoff worth of 0. 9, no length distinction re quirement and an first base pair match of 3 base pairs. Metagenome sequence information have been processed employing two fully automated open supply methods, the MG RAST v3. 0 pipeline plus the Fast Analysis of Multiple Metagen omes having a Clustering and Annotation Pipeline, offered through the Community Cyber infrastructure for Advanced Microbial Ecology Study and Examination. Taxonomic relationships be tween metagenomes had been analyzed by two complemen tary analyses using the MG RAST pipeline.
This 1st in depth gene catalogue rep resents a beneficial basel
This very first in depth gene catalogue rep resents a important baseline genomics resource for potential investigate into spider genetics and represents a to start with and basic phase in direction of understanding, and finally identifying, the genetic basis with the incredible shade poly morphism and patterning displayed by these animals. Solutions Samples, RNA extraction, normalization and sequencing Specimens of T. californicum have been collected from Albany Hill, Albany, Alameda County, California from beneath the leaves of blackberry plants during the early summertime when most folks are either adult or sub grownup. Specimens of T. grallator were collected from Lower Waikamoi Protect, Haleakala, East Maui, Hawaii from the undersides of leaves from the native Broussaisia arguta and Clermontia arborescens, as well as invasive ginger Hedychium gardnerianum.
All vital permits and permissions have been obtained and no more unique permissions had been essential for these species. To be able to facilitate the identification explanation of differentially expressed colour genes, two sets of animals were collected for every species. Just about every pool consisted of either the Yellow morph or maybe a mixture of Colored morphs. This very simple scheme is based mostly upon the fact that in all species studied, the Yellow morph seems to be recessive to all other colour morphs plus a equivalent scoring scheme is used previously, For T. californicum the Yellow pool comprised 20 Yellow folks as well as the Colored pool twenty individuals with the following morphs defined in Oxford. Red lines, Black spot, Black blob, White, Red ring A, Red ring B, Red stripe A, For T. grallator the Yellow pool consisted of 2 Yellow men and women and also the Colored pool 2 Red front and back men and women as defined in, All animals have been grownup females and consequently of a comparable size.
People have been examined to guarantee that no mites were current, starved for at least three days and then flash frozen at 80 C. Animals were homogenized and total RNA extracted using an RNeasy Mini Kit ac cording for the manufacturers guidelines. selleck chemical Five ug of total RNA was used to create an mRNA seq library from every sample pool. In addition, and as a way to recover the utmost quantity of genes, 2 ug of total RNA was con verted to cDNA utilizing a MINT cDNA synthesis kit and this was subsequently utilized to create a normalized cDNA library applying the TRIMMER kit, according to your producers instruc tions. Illumina sequencing libraries have been created from 50 ng of every normalized cDNA pool following the NEXTERA protocol and paired ends sequenced on either a Genome Analyzer II or Hi Seq 2000 sequencer, Sequence high quality assessment, pre processing and de novo assembly The raw sequence reads had been graphically inspected for top quality applying FastQC v.
While carboxyles terases and cytochrome P450s have important invo
Although carboxyles terases and cytochrome P450s have critical involvements in de toxification and sterol acquisition, cathespins, encapsulation proteins, mucin proteins, lipases, thaumatin domain proteins and lectins are hypothesized to perform funda psychological roles in mediating host microbe interactions. Not remarkably, GH 48 and GH 5 cellulases and GH 1 B glucosidases have been also remarkably expressed during the midgut, reflecting the nutritional importance of cellulose to A. glabripennis. GH 31 and GH 35 B galactosidases had been also really expressed, suggesting that galactan polymers current in hardwood hemicellulose or about the cell surface of microbes are also crucial sources of sugar for this in sect. Chitin deacetylase unigenes have been hugely abundant.
these enzymes can liberate acetate from insect or fungal chitin, which may be recycled for power or fatty acid production, Many different types of digestive proteinases were also extremely expressed and incorporated M16 peptidases, M14 carboxypeptidases, serine proteinases, and selleck inhibitor cysteine proteinases, which probably serve key roles in nitrogen extraction from plant or microbial cell wall proteins. Along with these digestive proteins, numerous unigenes predicted to encode hypothetical proteins with unknown functions had been abundant, suggesting that A. glabripennis encodes suites of novel proteins that may be appropriate for digestive physiology and improvement. Glycoside hydrolase profile comparisons As a result of comparisons of transcriptome libraries sam pled from a range of herbivorous insects, no key trends had been detected with regard to GH profiles and feeding habitats.
Euclidean distances involving insects that fed on equivalent substrates had been substantial in lots of circumstances and reflected robust differences in GH compositions. Thus, the gut transcriptome libraries didn’t demonstrate any substantial clustering trends by meals source, As an example, Agrilus planipennis and Dendroctonus ponderosae both feed in phloem and were observed in separate planes on the PCA ordination, recommended you read” indicating that there have been huge distinctions in GH household composition between these two insects. Likewise, A. glabripennis, Coptotermes formosanus, and Reticulitermes flavipes all feed in wood and had been also found in opposite quadrants in the PCA ordination. Although these insects are all capable of creating endogenous cellulases, A. glabri pennis creates various kinds of cellulases than the wood and phloem feeding insects compared in this research. As an example, the 2 termite species included within this examination predominately create GH 9 cellulases, whilst A.
gambiae was probably a refinement of that of An quadriannulatus
gambiae was probably a refinement of that of An. quadriannulatus. Additionally, this biased receptivity of An. gambiae antenna toward human derived odors may very well be even more augmented from the practical variations amongst orthologous ORs recommended by our sequence analyses. Future functional tests of AqOr odor tuning will even further enhance our understanding within this regard. Taken with each other, and offered the central part that ORs perform in defining host specificity, the anthropophagy of An. gambiae is most likely not derived from the evolution of any single OR precise for your objective of human host trying to find. Rather, we posit the receptivity bias inside the antenna of An. gambiae towards human host odors is possible the outcome on the cumulative effects of both practical divergences and changes in the abundance and distribution of widespread ORs currently current within the An.
gambiae species complex. Solutions Gene annotation The genome assemblies of An. gambiae and An. quadriannulatus had been downloaded from your internet sites of VectorBase and Broad Institute, respectively. The annotation of chemosensory genes was carried out following a past protocol, In brief, previously reported inhibitor PI-103 chemosensory genes from An. gambiae, Aedes aegypti, Culex quinquefasciatus, and D. melanogaster had been used as queries in TBLASTN searches towards the 2 anopheline genomes. Putative chemosensory gene coding loci were identified immediately after filtering out low scoring blast hits. For every locus, the query sequence that yield the highest bit score was selected as reference to execute homology primarily based gene prediction applying GeneWise, All gene models had been manually inspected and modified if needed.
All genomic data is available through VectorBase plus the annotated chemoreceptor sequences are listed in supplementary Table S1. Phylogenetic examination For every of your OR GR IR OBP households, protein sequences of genes inside the two mosquitoes had been aligned employing MAFFT, The numerous sequence alignments had been manually curated and poorly aligned regions have been eliminated applying trimAl with automated1 solution. recommended site Maximum probability trees had been constructed using RAxML plus the dependability of tree topology was evaluated with 100 bootstrap replicates. Resulting gene trees have been recon ciled together with the species phylogeny to estimate ancestral gene copy numbers and gene gain and reduction occasions. An orthologous group is defined being a very supported clade representing a single gene during the common ancestor of An.
gambiae and An. quadriannulatus. Evaluation of sequence divergence For every orthologous pair of chemosensory genes in An. gambiae and An. quadriannulatus, protein sequences were aligned applying MAFFT and also the corresponding nucleotide alignment was created utilizing a customized script, 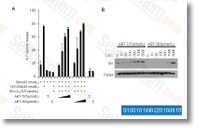 The fee of amino acid substitution and dN dS ratio have been calculated using PROTDIST and CodeML, respectively.
The fee of amino acid substitution and dN dS ratio have been calculated using PROTDIST and CodeML, respectively.
We then cross refer enced this for the down regulated gene set in
We then cross refer enced this towards the down regulated gene set in control versus muscle much less humeri, noting any genes enriched a lot more than 3 fold in mesenchyme compared to control humeri. these are indicated in column two of More file 1. Table S2. It can be potential that these genes are concerned in both cartilage and muscle improvement so no genes are already removed in the data set, even so, DE genes also displaying increased expression in mesenchyme com pared to regulate humeri needs to be taken care of with caution with respect to a skeletal exact response to mechanical stimu lation. Such genes have not been prioritised in any of our subsequent exploration of candidate mechanosensitive genes. The building humerus at TS23 constitutes distinctive cell and tissue populations at numerous stages of differen tiation like the joint region, the perichondrium plus the organised zones within the cartilage rudiment.
Thus the experimental layout employed here will capture genes related with different cells forms at dif ferent phases of differentiation. It can now be vital that you type out which cells and tissues have altered expres sion of certain genes. This could be selelck kinase inhibitor addressed to get a sub set of genes by in situ hybridisation, with an original ana lysis of 4 genes presented in Figure 6. It might be ad dressed within a large throughput manner by isolating exact cell populations employing laser microdissection from tissue sections, purification of RNA and quantitative RT PCR gene expression profiling, comparing management and mutant tissue from, one example is the hypertrophic, prehypertrophic or even the elbow joint re gion alone.
We implemented the two RNA sequencing and Micro array technologies in parallel to find out dif ferential expression. Microarray technological innovation has been utilised to find out expression of chondrogenic and osteogenic read this post here genes from establishing complete tissues, and from in vitro differentiation procedures, The use of RNA seq engineering to de scribe the transcriptome is more current, Previ ous direct comparisons amongst microarray and RNA sequencing based approaches to reveal alterations in gene expression amongst tissues reported that RNA seq identified a lot more DE genes, We also noticed that RNA seq is additional delicate in reproducibly detecting alterations in gene expression, detecting far more genes al tered at decrease quantitative amounts, This was additional emphasised by minimizing the stringency from the statistical analysis to p 0.
08, which increased the amount of genes detected by micro array particularly, An illustration from the im portance from the elevated sensitivity and reproducibility of RNA seq is proven from the Spp1 gene which did not display statistical significance by microarray but is verified by qRT PCR and in situ hybridisation, The larger dynamic assortment and greater reproducibility across replicates has also been found in other scientific studies.
Pharmacophore primarily based screening to identify anti cancer l
Pharmacophore based screening to recognize anti cancer leads On screening the compound library primarily based to the phara macophoric hypothesis DDHRR. eight, the resulting 7409 compounds had been subjected to XP docking towards cathe psin L. The resultant six compounds that scored over the threshold have been selected. The 3D QSAR model formulated making use of precisely the same congeneric series as that on the pharamacophoric model was utilized to predict the exercise within the resultant 6 compounds. The docking scores and predicted pursuits are summarised in Table four. We report the leading two scor ing compounds obtained and evaluated through this mixed approach. The XP score provides the extent of binding affinity on the respective lead molecules with Cathepsin L, all of them lying under the specified threshold.
We focused about the catalytic triad comprising of residues Cys25, Met161 and Asp162 and analyzed the interactions taking place among Cathepsin L and the thiosemicarbazone series, The initial compound reported selleck chemicals Wnt-C59 is usually a bulky ringed structure that interacted together with the catalytic triad along with other residues. Ala138, Gly139, Trp26, Gly68, Ala135, Gly164, Leu69, and Ala214, The subsequent leading scoring candidate four hydroxy 1 methyl quinolin 2 one], once more a bulkier 1, weakly interacted using the catalytic triad aside from Gly67, Gly68, Leu69, Met70, His163, Ala135 and Ala214, It may possibly be inferred that due to the steric hindrance brought on by its bulky aromatic groups, APQ fails to interact closely with Cys25, Met161 and Asp162. The align score refers towards the extent of similarity with all the picked hypothesis, DDHRR. eight.
Align score was identified to become highest for NFP being 1. 195091 when for APQ it had been 0. 974276. selleck KU-0060648 We predicted the routines of the top scoring compounds using the generated 3D QSAR model. The high predicted pursuits of NFP and APQ advised that it is really worth to take into consideration them potent cathepsin L inhibitors. Conclusion We implemented a combined approach to screen potent cathepsin L inhibitors that promised to emerge as necessary leads in cancer investigate owing for the part that cathepsin L plays throughout tumor growth and metastasis. A congeneric set belonging towards the thiosemicarbazone class of molecules which are regarded to inhibit human cathepsin L was cho sen to construct a 3D QSAR model as well as a pharmacophore model.
The former related the structure within the molecule with its exercise quantitatively although validating the relation ship employing statistical parameters whereas the later on pointed out the minimal structural options crucial for any molecule for its biological activity 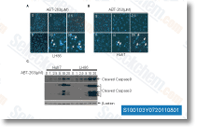 as well as provided an insight into the mode of binding together with the target. Making use of these two approaches of ligand based drug developing we screened a chemical library primarily based to the pharamacophoric hypothesis after which predicted their action applying the 3D QSAR model.
as well as provided an insight into the mode of binding together with the target. Making use of these two approaches of ligand based drug developing we screened a chemical library primarily based to the pharamacophoric hypothesis after which predicted their action applying the 3D QSAR model.
Inhibitors for ERK1 2 and its con trol substance, p38, PI3K, Akt
Inhibitors for ERK1 2 and its con trol substance, p38, PI3K, Akt were from Calbiochem, Rabbit polyclonal p38 MAPK and Akt, and monoclonal phos pho p38 MAPK, phospho Akt, p44 p42 MAPK, and phospho p44 p42 MAPK antibodies were obtained from Cell Signaling Technologies, Inc, Mouse monoclonal GAPDH, used as western blot control for equal gel load of protein, was purchased from Santa Cruz Biotechnol ogy Inc, As assessed from the Limulus Amebocyte Lysate Pyrotell, the thrombin and trypsin preparations in our study contained less than 0. 03 EU LPS ml. RNA isolation, reverse transcription and Quantitative RT PCR Single stranded cDNA was synthesized from total RNA and utilized to carry out QRT PCR with gene specific pri mers as described previously, Unstimulated HOKs and samples with no RT served as negative controls.
A melting curve was carried out on the end of each QRT PCR to make certain the gene solution was particular. Gly ceraldeyde 3 phosphate selleck chemical dehydrogenase was utilised as the selected housekeeping gene for normaliza tion. The primer sequences for GAPDH, CXCL5, CXCL3 and CCL20 have been described previously, Data had been analyzed working with Pfaffl process to cal culate fold modify of gene expression, Measurement of chemokines in culture supernatant Secreted CXCL5 and CCL20 have been measured in culture supernatant employing Duoset ELISA kit, The absolute concentration in just about every sample was calculated according to sample yielding optical density employing the regular curve. Chemokine secretion from every condition was normalized to chemokine made at baseline level in unstimulated management group, and benefits are presented as % of unstimulated management cells.
ELISA assay for quantification of p38 and ERK1 two phosphorylation To assess phosphorylation of p38 and ERK1 2, handled cells had been homogenized in lysis buffer in accordance together with the suppliers protocol, Immediately before adding to cells, protease and phosphatase inhi bitors, PMFS and sodium orthovanadate, Halt Protease Inhibitor Cocktail and Halt selleck chemical SAR302503 Phosphatase Inhibitor Cocktail, had been extra for the lysis buffer. Phosphorylation of p38 and p42 44 was assessed in cell lysate working with Duoset antibody pairs following manufacturers protocol, Total p38 was quantified by a similar strategy and served for normalization. Complete p38 was proportional to complete protein measured in each and every sam ple and also to the amount of cells plated in every nicely.
Western blot examination Total and phosphorylated kinase expression in untreated and agonist stimulated HOK cells was determined by Western immunoblot evaluation. Samples of complete cell lysate supernatant proteins, solubilized in 1X NuPage LSD sample and lowering buffer, had been resolved conco mitantly with Precision Plus Dual Shade protein molecu lar excess weight standard in NuPAGE four 12% Bis Tris mini gel and MOPS SDS working buffer in XCell SureLock Mini Cell, trans ferred to PVDF membrane in XCell II Blot Module making use of electrophor esis programs and protocols of InVitrogen Corp, PVDF membranes have been blocked for 1 h with 1% BSA, 1%PVP and 1% PEG in wash buffer, then.
Inhibitors for ERK1 two and its con trol substance, p38, PI3K, Ak
Inhibitors for ERK1 two and its con trol substance, p38, PI3K, Akt were from Calbiochem, Rabbit polyclonal p38 MAPK and Akt, and monoclonal phos pho p38 MAPK, phospho Akt, p44 p42 MAPK, and phospho p44 p42 MAPK antibodies had been obtained from Cell Signaling Technological innovation, Inc, Mouse monoclonal GAPDH, utilised as western blot handle for equal gel load of protein, was bought from Santa Cruz Biotechnol ogy Inc, As assessed through the Limulus Amebocyte Lysate Pyrotell, the thrombin and trypsin preparations in our examine contained less than 0. 03 EU LPS ml. RNA isolation, reverse transcription and Quantitative RT PCR Single stranded cDNA was synthesized from complete RNA and used to perform QRT PCR with gene specific pri mers as described previously, Unstimulated HOKs and samples with out RT served as adverse controls.
A melting curve was carried out at the end of each QRT PCR to be sure the gene solution was unique. Gly ceraldeyde 3 phosphate order Dabrafenib dehydrogenase was made use of since the chosen housekeeping gene for normaliza tion. The primer sequences for GAPDH, CXCL5, CXCL3 and CCL20 are already described previously, Data have been analyzed working with Pfaffl process to cal culate fold transform of gene expression, Measurement of chemokines in culture supernatant Secreted CXCL5 and CCL20 have been measured in culture supernatant employing Duoset ELISA kit, The absolute concentration in each and every sample was calculated determined by sample yielding optical density utilizing the typical curve. Chemokine secretion from each and every affliction was normalized to chemokine generated at baseline degree in unstimulated handle group, and outcomes are presented as % of unstimulated manage cells.
ELISA assay for quantification of p38 and ERK1 2 phosphorylation To assess phosphorylation of p38 and ERK1 two, taken care of cells were homogenized in lysis buffer in accordance together with the makers protocol, Immediately prior to incorporating to cells, protease and phosphatase inhi bitors, PMFS and sodium orthovanadate, Halt Protease Inhibitor Cocktail and Halt read what he said Phosphatase Inhibitor Cocktail, have been additional towards the lysis buffer. Phosphorylation of p38 and p42 44 was assessed in cell lysate applying Duoset antibody pairs following producers protocol, Complete p38 was quantified by a very similar approach and served for normalization. Complete p38 was proportional to complete protein measured in every single sam ple and to the quantity of cells plated in just about every properly.
Western blot analysis Total and phosphorylated kinase expression in untreated and agonist stimulated HOK cells was determined by Western immunoblot examination. Samples of entire cell lysate supernatant proteins, solubilized in 1X NuPage LSD sample and lowering buffer, had been resolved conco mitantly with Precision Plus Dual Colour protein molecu lar bodyweight conventional in NuPAGE 4 12% Bis Tris mini gel and MOPS SDS operating buffer in XCell SureLock Mini Cell, trans ferred to PVDF membrane in XCell II Blot Module employing electrophor esis programs and protocols of InVitrogen Corp, PVDF membranes have been blocked for 1 h with 1% BSA, 1%PVP and 1% PEG in wash buffer, then.
